The main motivation behind my research is coming up with new and better methods for testing drugs and other substances to see if they are effective and safe. I've written previously about some of the promising animal testing alternatives that are being created.
Last week I went to a workshop run by the national centre for the replacement/refinement/reduction of animals in research (NC3Rs) and POEMS. Biologists brought 3Rs problems, and a group of computational and mathematical modellers tried to solve them.
There were five different projects presented. One was about using fruit flies to model diseases like Alzheimer's and Parkinson's by seeing how their courting behaviour is affected, another was modelling the way that nerves respond to bladder filling, one was related to immunoglobulin, and another was modelling retina damage in shaken baby syndrome (so much respect for the people who picked that project. It's really important but heartbreaking). The project I chose was about testing for neurodevelopmental safety.
The problem
If you affect the amount of thyroid hormones in the body of a pregnant woman, it can have huge effects on the developing foetus. A lot of potential drugs, potential industrial chemicals and potential pesticides turn out to significantly reduce the amount of thyroid hormones in the blood, meaning that they aren't safe to use around people.
Currently if you want to bring a new chemical to market, you have to test for thyroid effects using about 40 pregnant rats, each of which could be pregnant with up to 10 pups. We wanted to see if it was possible to use data from earlier studies on a few adult male rats to predict these effects, without having to use any extra animals. This method would still use data from animal tests, but would reduce the number of animals used, and could pave the way towards a non-animal method later on.
The picture below shows a simplified version of the pathway that thyroid hormones take through the body. T4 is produced in the mother's thyroid gland and then secreted into her blood. From here, it can either be removed by the liver, or carried into the blood of her foetus. It then travels from the foetus' blood to its developing brain cells. Inside the brain cells, T4 is converted into T3, which is essential for the brain to develop correctly (for a review, see Williams, 2008).

The process
First, we all sat down together and came up with a set of equations to describe the way that all the components of the regulatory network interacted with each other. We used simple ordinary differential equations, which you might recognise from chemistry lessons.
For example, if a molecule A turns into a different molecule B at a rate of c molecules per second, and B turns into A at a rate of d molecules per second, you can say that
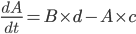
and
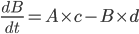
This way, given starting concentrations for A and B, you can work out how many molecules of each you have at a given time. We used this method to write down equations to describe the whole network.
After this, we each followed a different path of investigation. One researcher tried to derive analytically an equation to describe what steady state the system will reach, while others performed sensitivity analysis and parameter fitting. I was in a group that were looking at Petri nets, which are a different way of describing reaction networks, with some useful tools for analysis.
An animation of the petri net for this system is shown below. The circles (known as "places"), represent possible states that molecules can be in, the numbers inside the places represent the number of molecules in that state. The squares are transitions - usually representing some kind of reaction or binding event - and the red dots moving between places and transitions are the molecules as they change from place to place.
Petri nets seem like a really useful way of modelling biological systems for people without much experience of ODEs or programming, because they are a direct analogue for a system of ODEs with all the rate constants set to 1. The next stage in our analysis will be to parameterise this model, and use some graph theory tools to find out new things about the system. I haven't done graph theory since A-level, so this should be quite interesting!
