As I mentioned in my previous post about my adventures in FLAME modelling, I started out with code from my lecturer. I started with three two-dimensional models, each of which contained a different type (genus) of bacteria (Vibrio, Sar11 and "Bacteria X"). Each bacteria agent had X and Y co-ordinates, an ID and a radius. The radius was different for each type of bacteria, and it defined the rules that applied to that bacteria. They moved around by Brownian motion, and if they hit a virus there was a certain probability that they would be killed. The phages moved around in a similar fashion.
My code can be downloaded from here. In this post, I'm going to take you through how I converted these three models into a single model, added a third dimension, and persuaded the viruses and bacteria to replicate. This is a tale of how it's possible to do some quite cool things to a program even if you don't fully understand how it works.
If you're particularly interested, you can download a PDF of the report I handed in here. It starts with a brief review of a mathematical bacteriophage model, then goes on to the FLAME model in the second half (page 2). I got a mark of 95% for it, so I'm showing it off to everyone ^_^
Three becomes one.
This part was arguably the simplest. The C file phageAndBacteriaV6.c contained conditional statements like this one:
if (BACTERIARADIUS == 100){
//SAR11
yUpperBorder = 24;
yLowerBorder = 736;
} else if (BACTERIARADIUS == 500){
//Bacteria X
yUpperBorder = 40;
yLowerBorder = 720;
} else {
//Vibrio
yUpperBorder = 57;
yLowerBorder = 703;
}
Which handily deals with all the different behaviours of the bacterial species. So all I had to do was combine the bacteria from all 3 versions of the 0.xml file together, and change their IDs to avoid overlap. Then I used conditional display in the FLAME visualiser to make them different sizes and colours in my animation.
Going from 2D to 3D
This was slightly harder - but nowhere near as difficult as I'd anticipated. Broadly, I found everywhere that X and Y were referenced in the code, and added a Z. I replicated a couple of position-checking functions wholesale (including the one above) and added some variables in some places. I found it useful to use "grep -i", followed by the name of the thing I'd changed, at the command line to look for where variables were used.
You're probably familiar with the equation to find the length between 2 points in 2 dimensions (from good old Pythagoras):
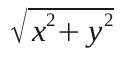
Extending it to three dimension gives:
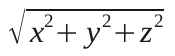
So I changed that function, and that does it for phageAndBacteriaV6.c.
I used the FLAME editor to change phageAndBacteriaV6.xml to include the variable “bacteriaZ” as a double belonging to the "Bacteria" agent and to the "bacteriaInformation" message. I also added the variable “phageZ” as a double to the Phage agent and the phageInformation message.
Next, I needed to add starting Z positions for the bacteria and viruses to 0.xml. This was a bit more tricky. I started out by simply using "Find and Replace" in Gedit to change:
</bacteriaY>
to
</bacteriaY>
<bacteriaZ>120</bacteriaZ>
and the same with phageY and phageZ. This was great, because I could then try compiling the model, by running xparser, then "make". There was only one error on the first attempt (I'd forgotten an asterisk somewhere) and then it all worked fine. I opened up the files using the FLAME visualiser and added the Z component to the visualisation, and there they were, bobbing around.
Great! Except that the bacteria and phages all started from the same Z position (i.e. in the same plane). I wanted them in random places, so I made up a little bash script:
#! /bin/bash
RANGE=780
for i in {1..30}
do
echo $i
number=$RANDOM
let "number %= $RANGE"
let "number=$number + 57"
sed -ie "0,/<phageZ>120/{s/<phageZ>120/<phageZ>$number/}" ./test.xml
done
Which I ran once, then changed the word "phage" to "bacteria" and ran again. This distributed them randomly within the boundaries of my little universe (between 57 and 837 units, somewhat arbitrarily).
Bacterial replication
I was only supposed to be adding phage replication, but I thought bacterial would be interesting (and straightforward), so I did that first. I have to admit I was stumped to begin with, but upon reading the manual (which I should definitely have done first), I discovered the built-in function add_agentname_agent(var1,...,varN). So, based on a random number, I can get my bacteria to divide occasionally. I included this bit of code inside the bacteriaRandomMove() function, which is called at every time step.
double diceRoll = 0;
diceRoll = ((double)rand()/((double)RAND_MAX)); //0 and 1
if (diceRoll < = dividingProbability) { printf("Bacteria %d at bx %f by %f bz %f has divided \n", BACTERIAID, BACTERIAX, BACTERIAY, BACTERIAZ); add_Bacteria_agent(newb_ID, BACTERIAX, BACTERIAY, BACTERIAZ, BACTERIARADIUS, DEFENSEFACTOR); }
Easy peasy! The only problem with this, is that the bacteria ends up with the same ID as its mother. This means that if the one bacteria is killed, its descendants will be as well. According to a trusted source (my awesome friend Gavin), this is an outstanding bug in FLAME, as there is no way to give consecutive numbers to new agents. People often solve this problem by giving agents random IDs over a large range - which does run the risk of occasional collisions, but is certainly better than my method.
Phage burst
There was already a function that kills a bacteria when it is hit by a successful phage attack, so I just had to add in my phage replication code there.
int makeNewPhage(int lphageID, double lphageLife, double lphageX, double lphageY, double lphageZ, double lphageRadius, int lsetSeed, double bactRadius) {
int burstSize = 0;
if (bactRadius == 100){
//SAR11
burstSize = 1;
} else if (bactRadius == 500){
//Bacteria X
burstSize = 5;
} else {
//Vibrio
burstSize = 9;
}
printf("%d\n",burstSize);
int i;
for(i=0;i<burstSize;i++) { add_Phage_agent(lphageID, lphageLife, lphageX, lphageY, lphageZ, lphageRadius, lsetSeed);}
return 0;
}
